Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Nature Communications
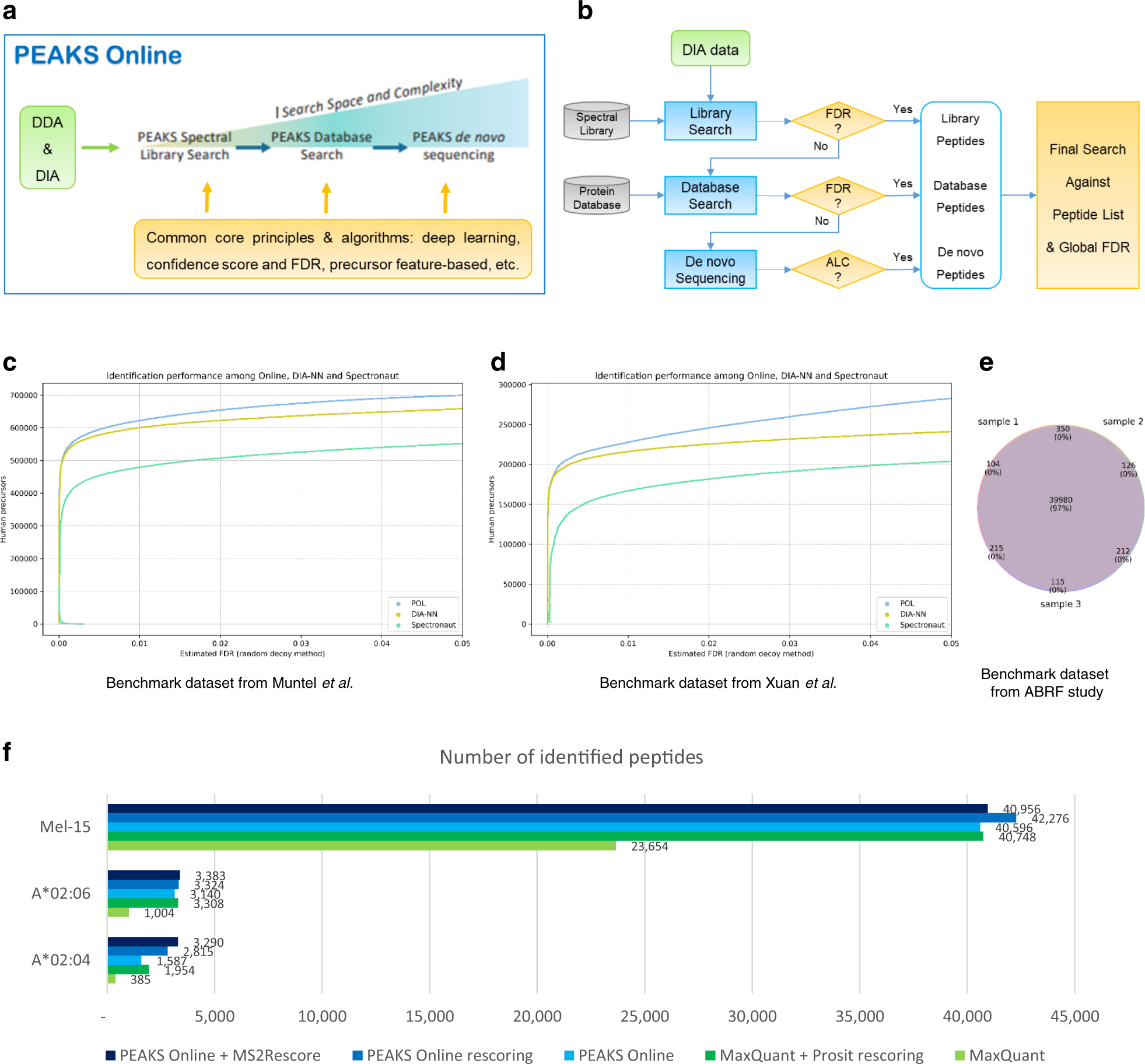
A streamlined platform for analyzing tera-scale DDA and DIA mass spectrometry data enables highly sensitive immunopeptidomics | Nature Communications
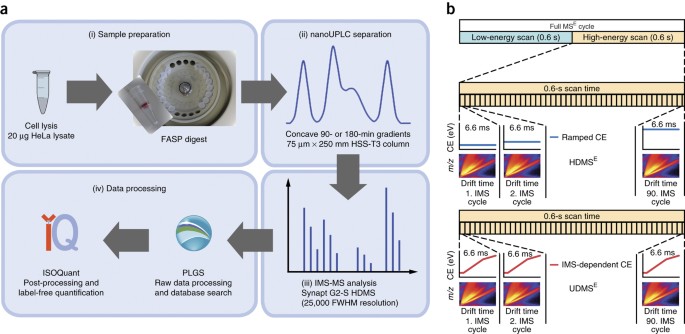
Label-free quantification in ion mobility–enhanced data-independent acquisition proteomics | Nature Protocols

Sensitive Immunopeptidomics by Leveraging Available Large-Scale Multi-HLA Spectral Libraries, Data-Independent Acquisition, and MS/MS Prediction - ScienceDirect

Technical comparison of DDA and different types of DIA in a biological... | Download Scientific Diagram
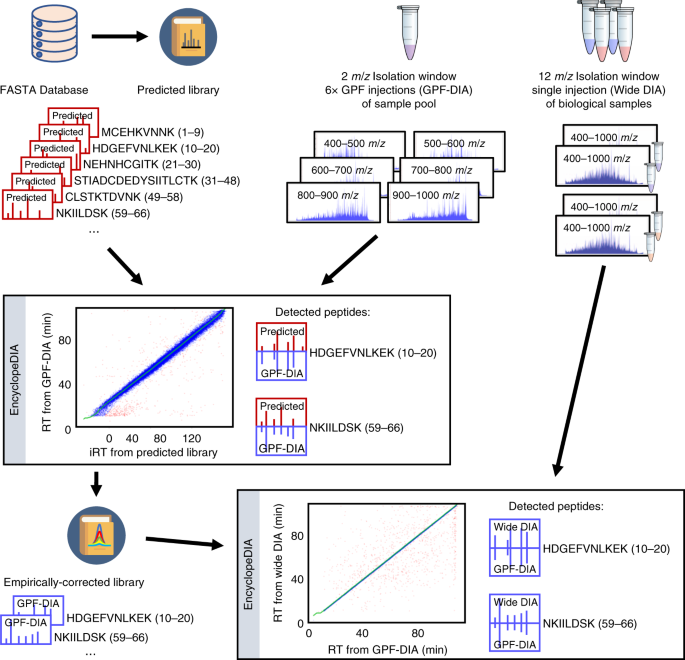
Generating high quality libraries for DIA MS with empirically corrected peptide predictions | Nature Communications
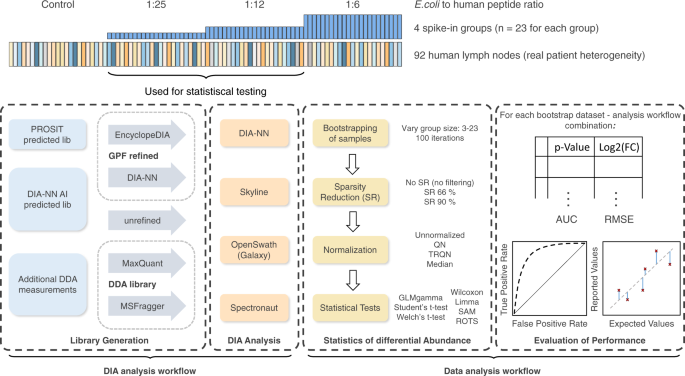
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity | Nature Communications
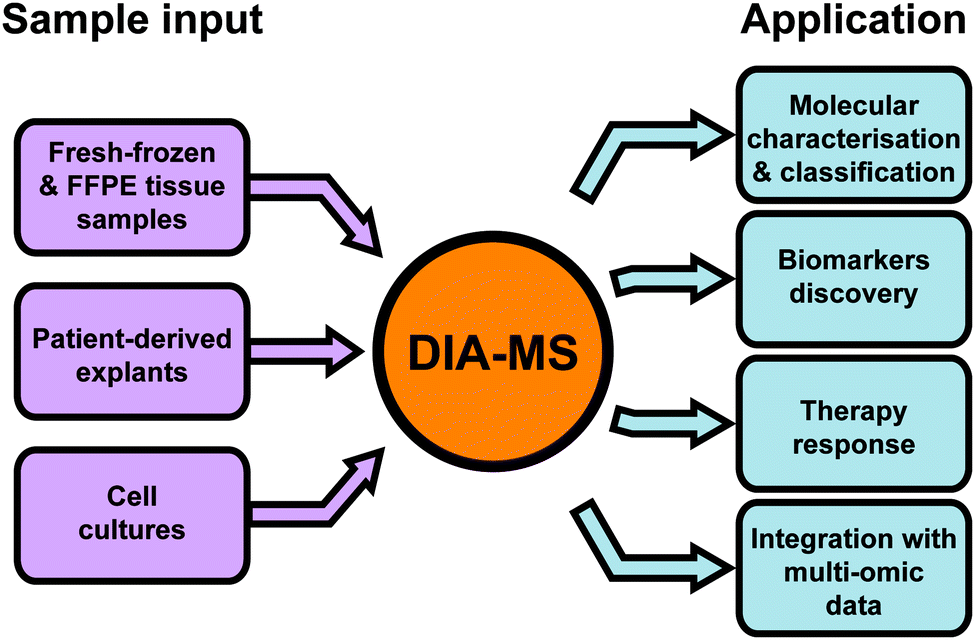
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
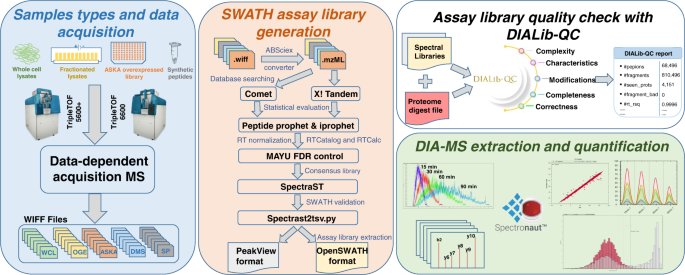
A comprehensive spectral assay library to quantify the Escherichia coli proteome by DIA/SWATH-MS | Scientific Data
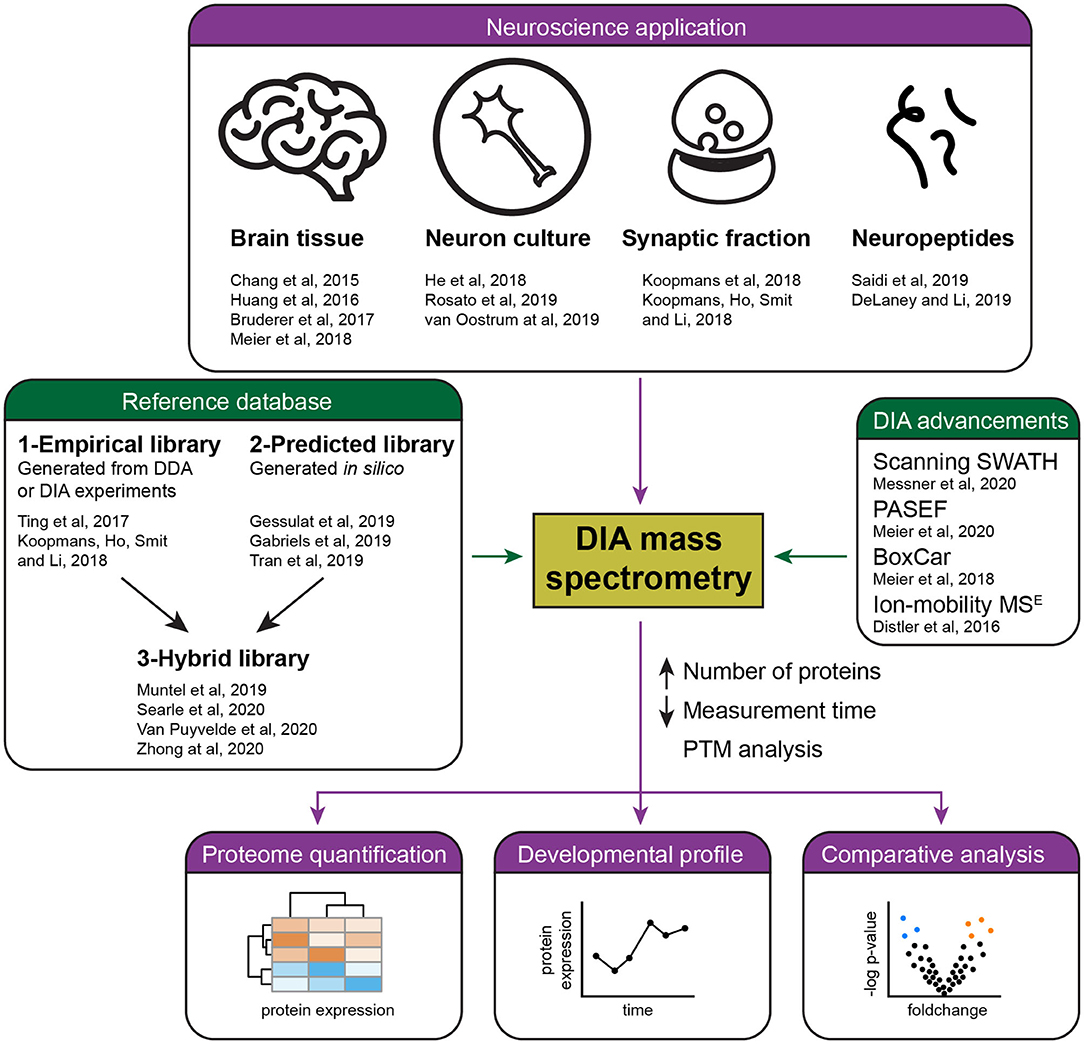
Frontiers | Recent Developments in Data Independent Acquisition (DIA) Mass Spectrometry: Application of Quantitative Analysis of the Brain Proteome
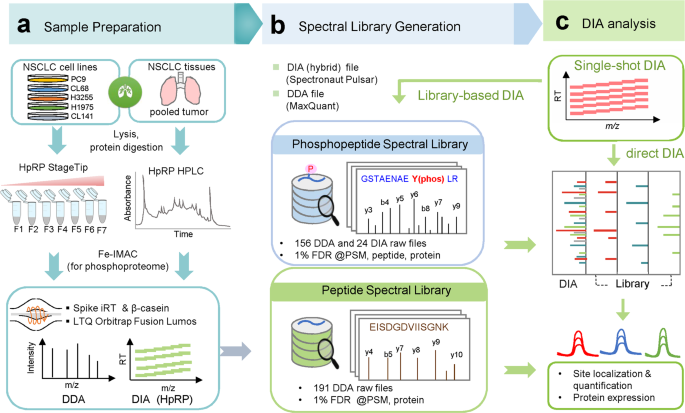
A data-independent acquisition-based global phosphoproteomics system enables deep profiling | Nature Communications
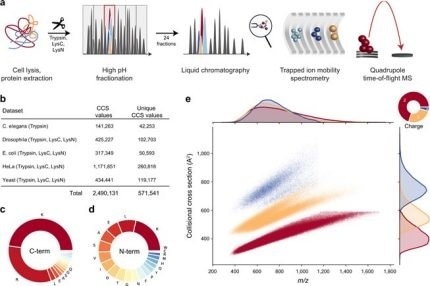
New Nature Communications publication by Mann & Theis Groups harnesses the benefits of large-scale peptide collisional cross section (CCS) measurements and deep learning for 4D-proteomics

Deep Proteomics Using Two Dimensional Data Independent Acquisition Mass Spectrometry | Analytical Chemistry

DIAproteomics: A Multifunctional Data Analysis Pipeline for Data-Independent Acquisition Proteomics and Peptidomics | Journal of Proteome Research
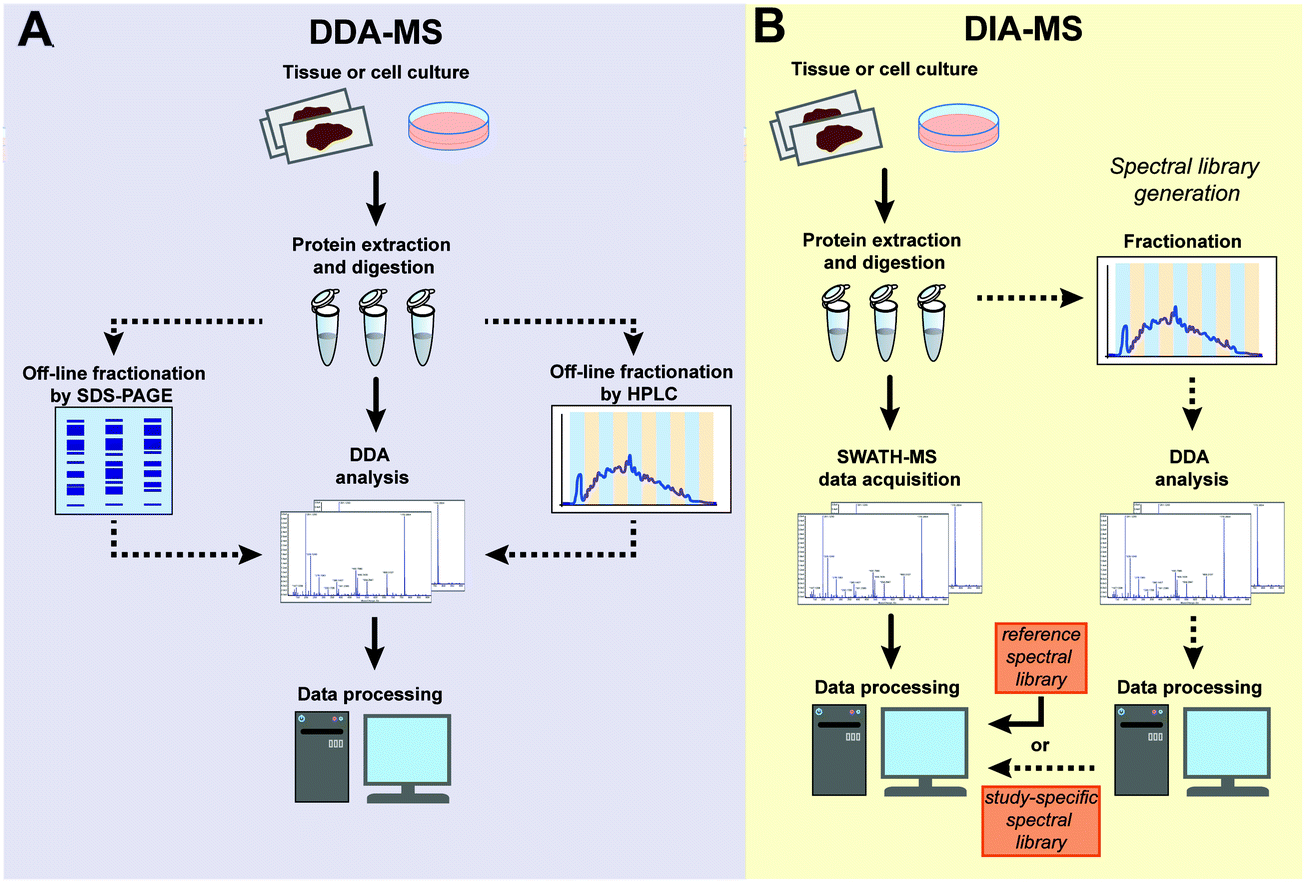
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H

Direct data-independent acquisition (direct DIA) enables substantially improved label-free quantitative proteomics in Arabidopsis | bioRxiv
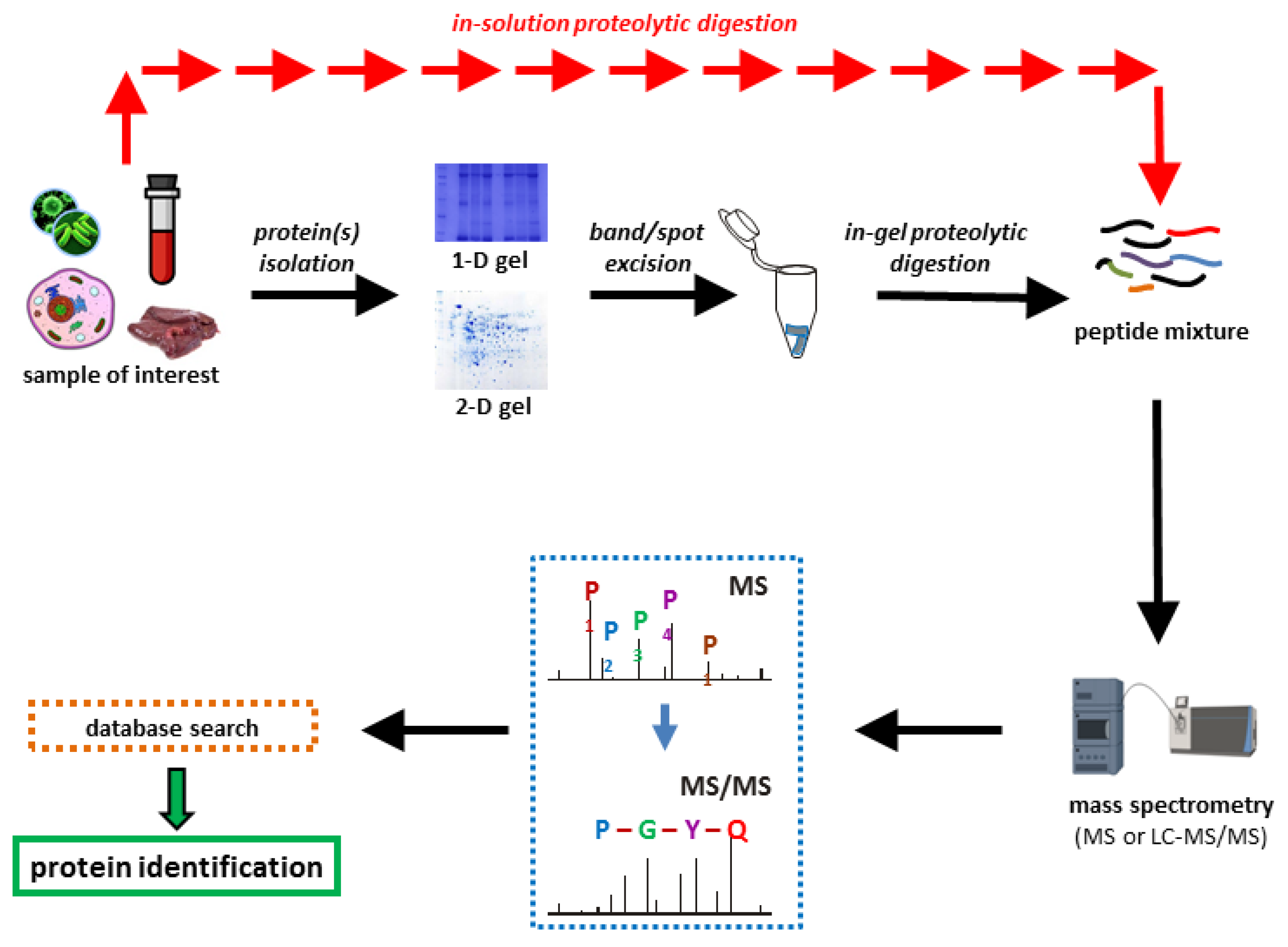
Proteomes | Free Full-Text | A Critical Review of Bottom-Up Proteomics: The Good, the Bad, and the Future of This Field
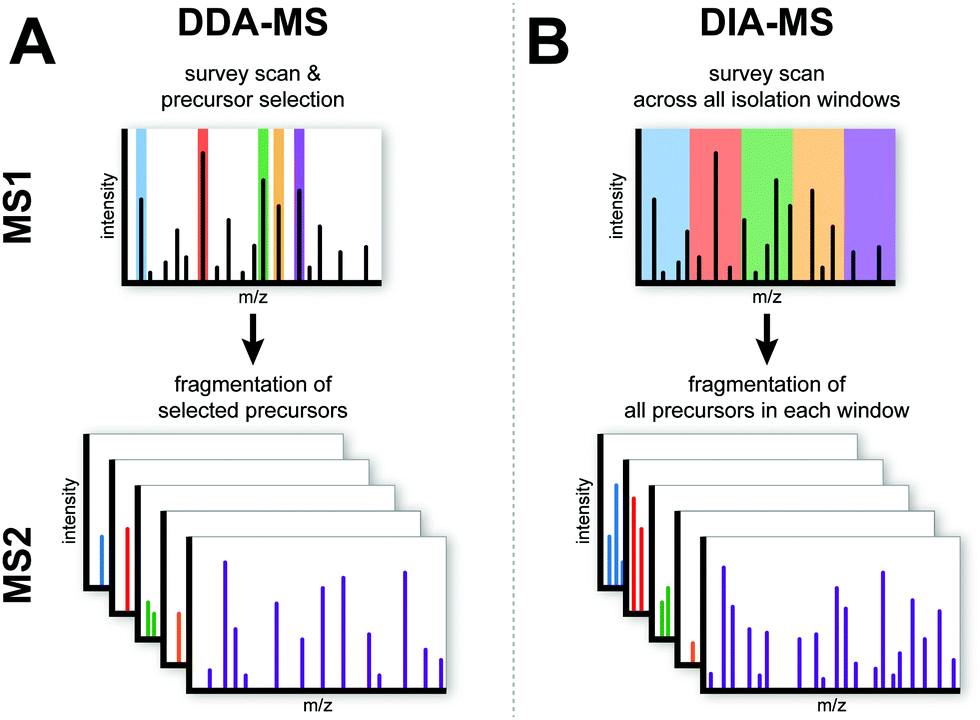
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H

Group-DIA: analyzing multiple data-independent acquisition mass spectrometry data files | Nature Methods

Variability in Mass Spectrometry-based Quantification of Clinically Relevant Drug Transporters and Drug Metabolizing Enzymes | Molecular Pharmaceutics
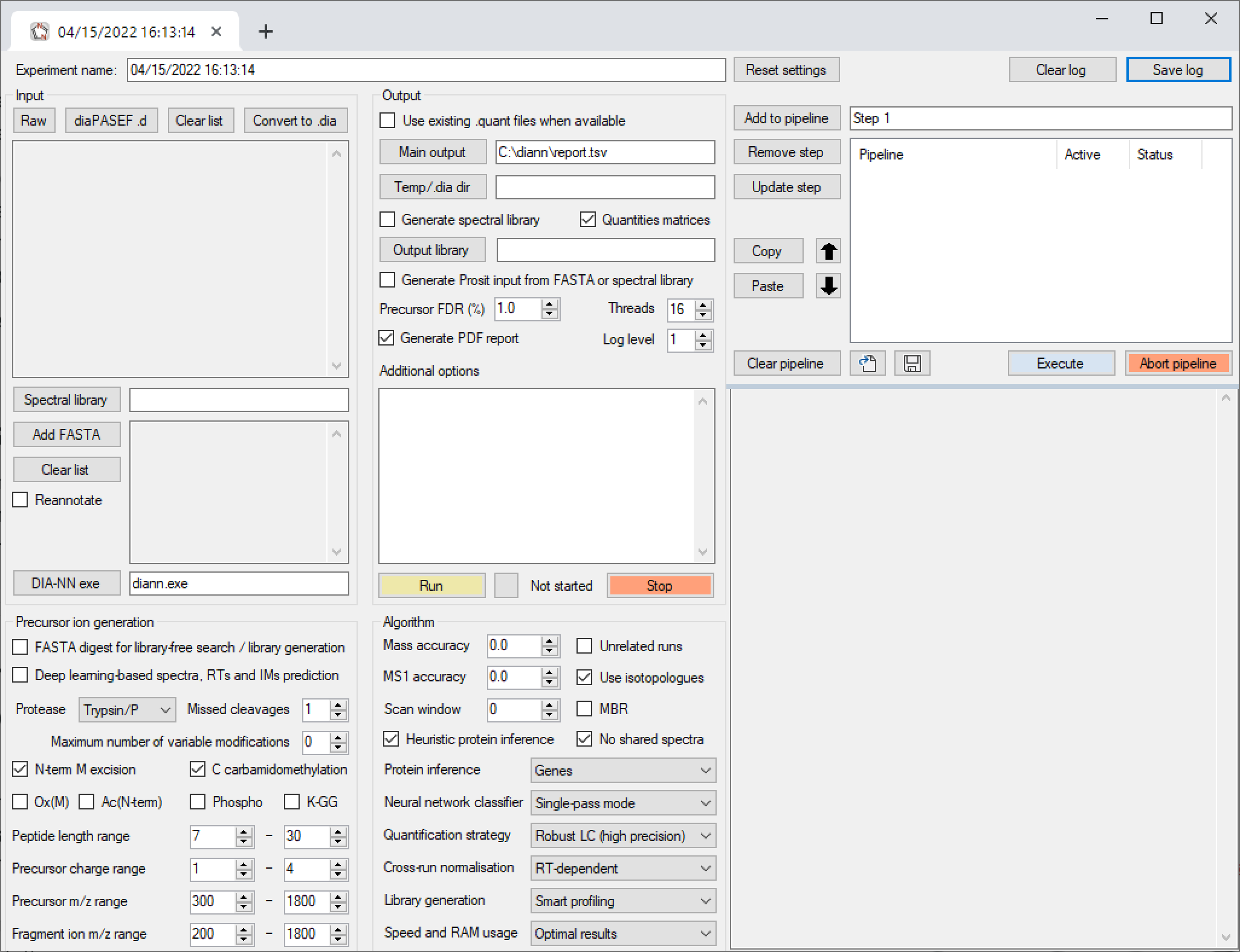
GitHub - vdemichev/DiaNN: DIA-NN - a universal automated software suite for DIA proteomics data analysis.

Benchmarking Bioinformatics Pipelines in Data-Independent Acquisition Mass Spectrometry for Immunopeptidomics - ScienceDirect