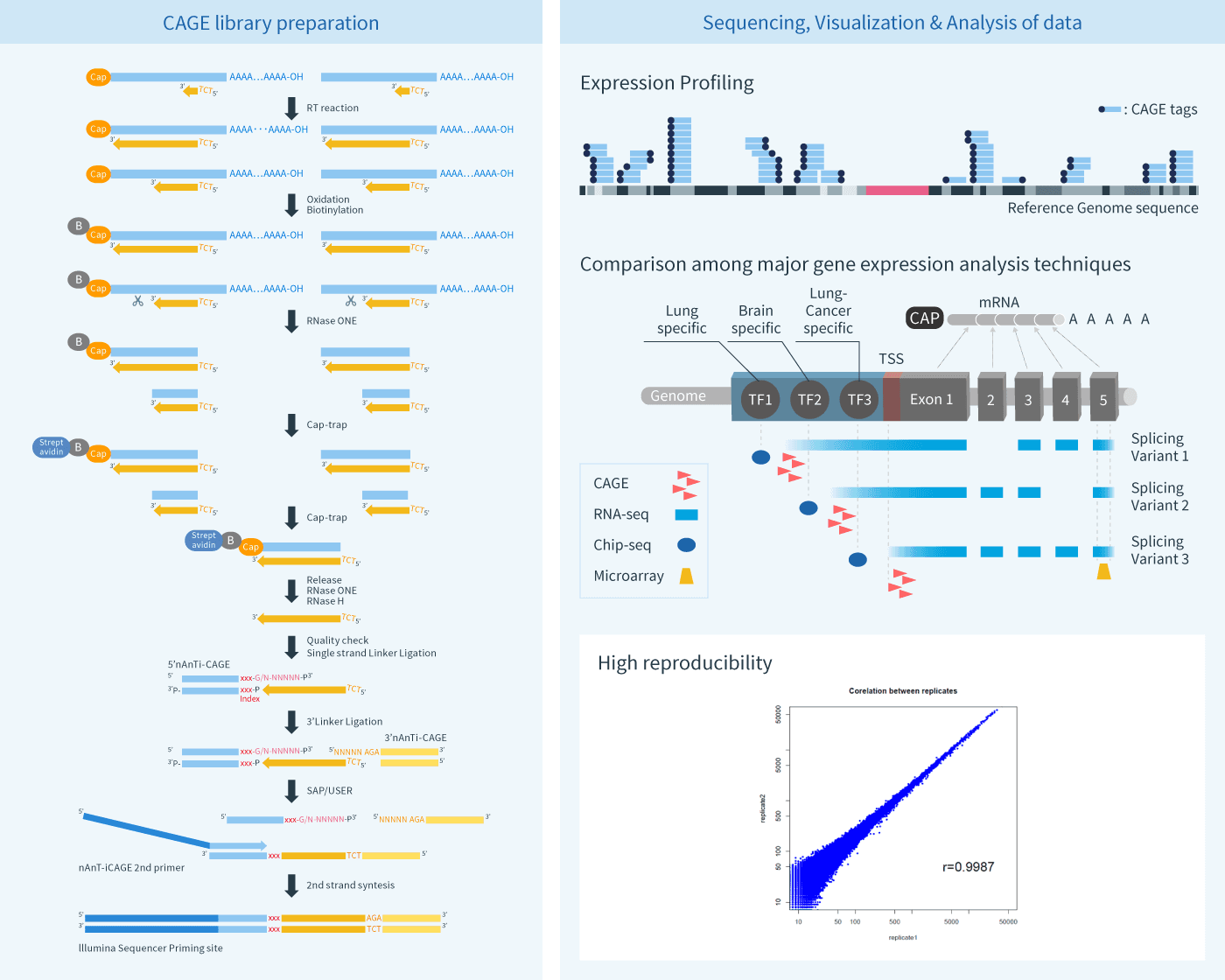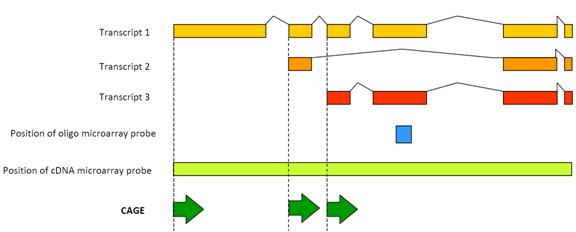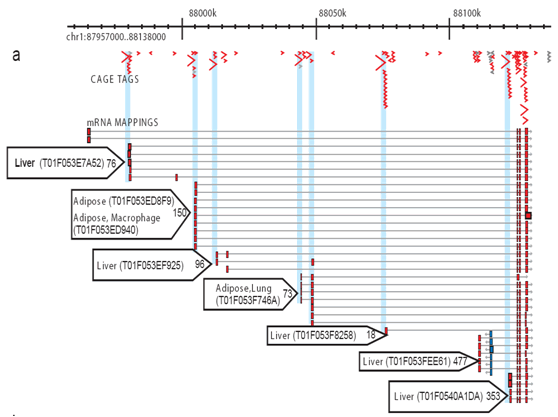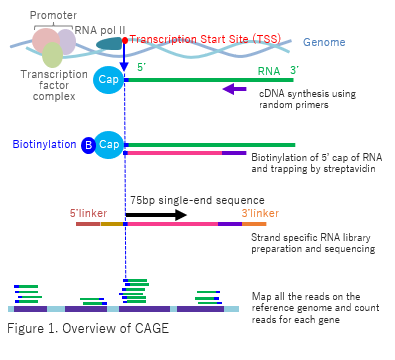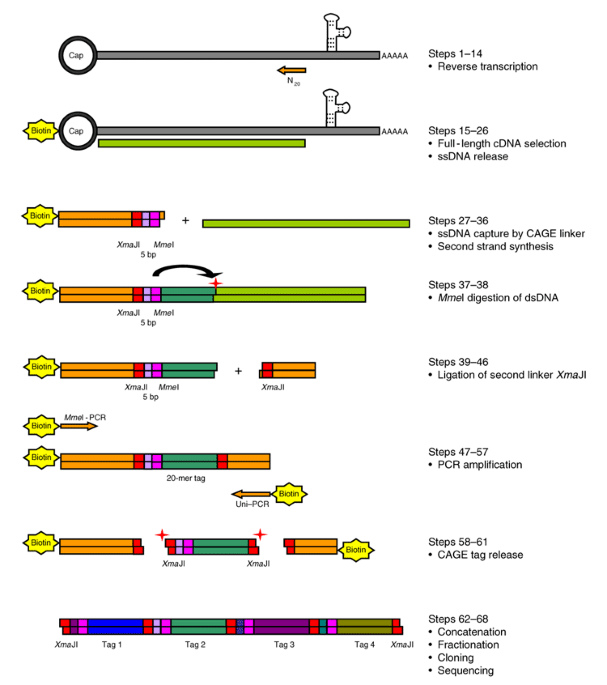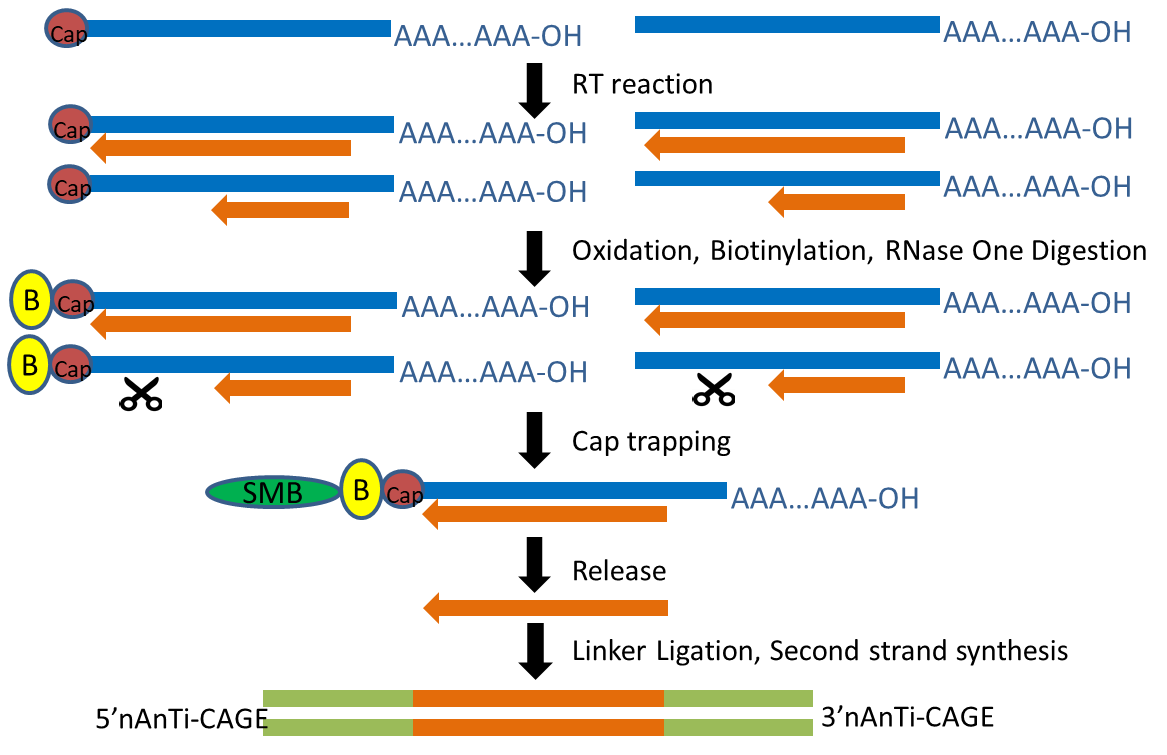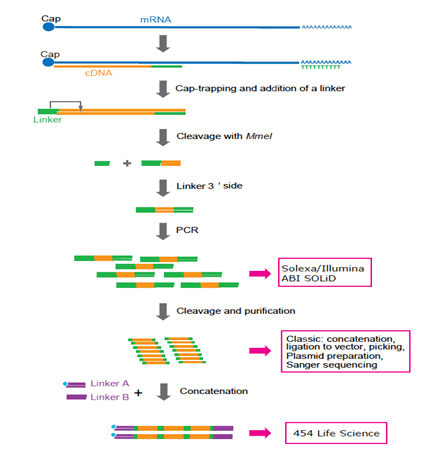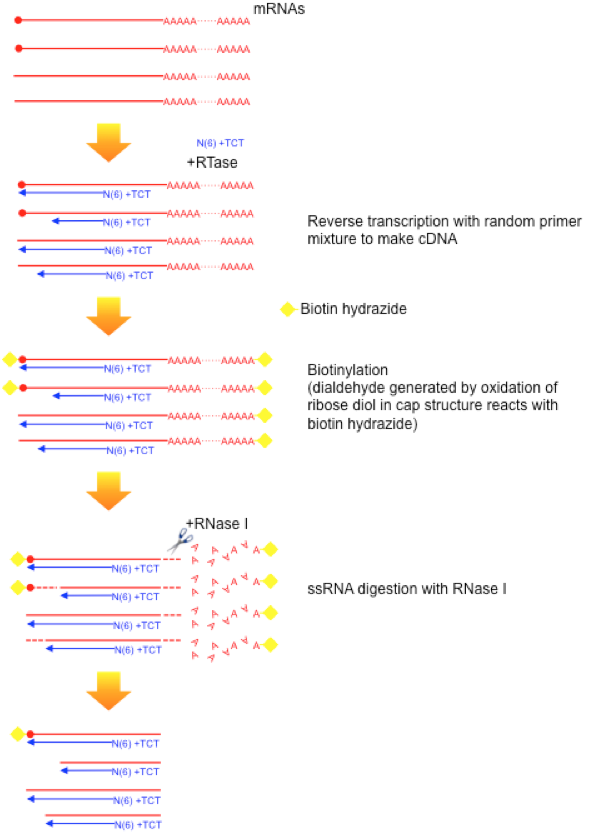
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター

Schematic diagram of CAGE and RNA-Seq read coverage for a gene with two... | Download Scientific Diagram

Cap analysis of gene expression (CAGE) sequencing reveals alternative promoter usage in complex disease | bioRxiv

CAGE-seq characterization of alternative promoter usage and full-length... | Download Scientific Diagram
CAGE-seq analysis of Epstein-Barr virus lytic gene transcription: 3 kinetic classes from 2 mechanisms | PLOS Pathogens
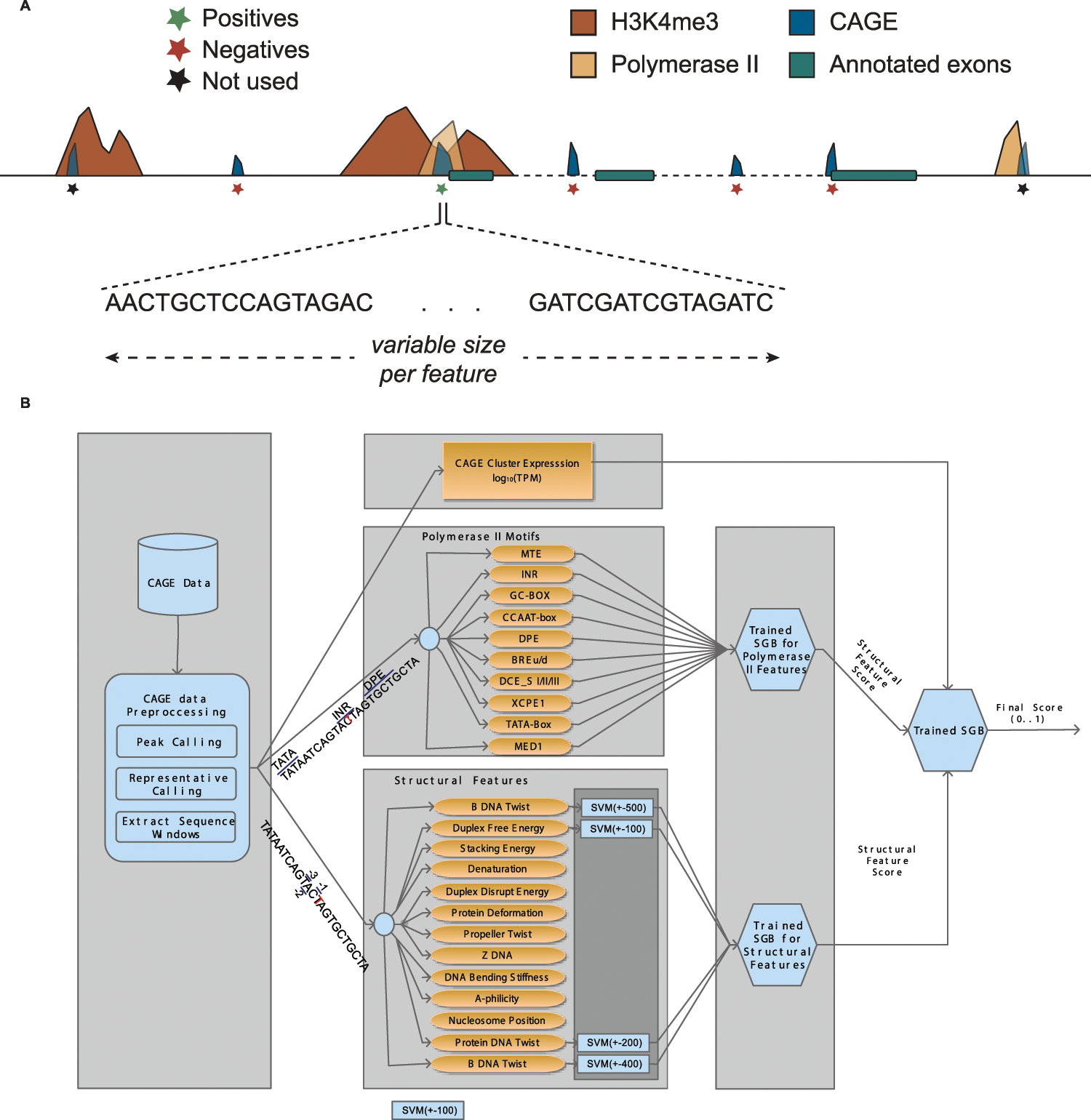
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports
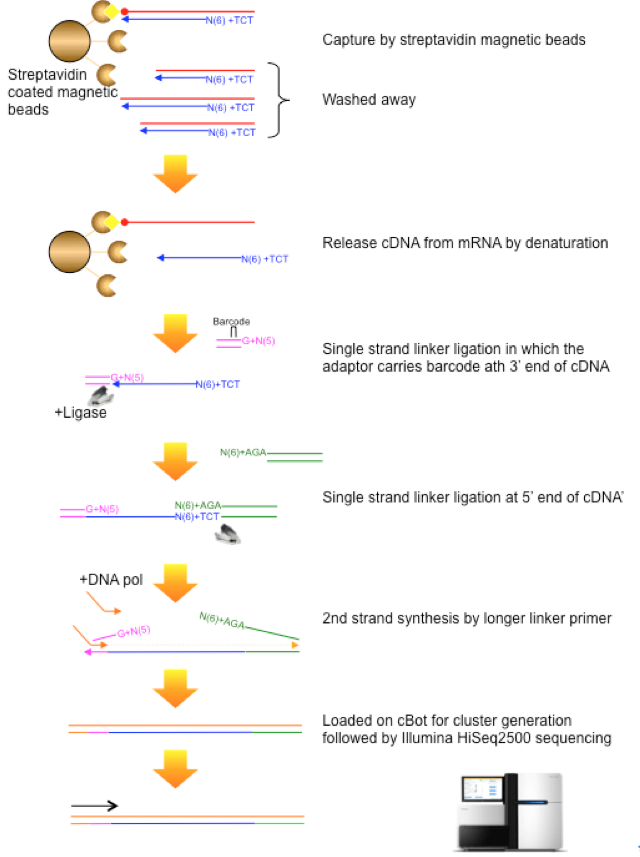
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター